Cardiac Image Registration
Jasbir Sra
Electrophysiology Laboratories, Aurora Sinai/Aurora St. Luke’s Medical Centers,University of Wisconsin School of Medicine and Public Health-Milwaukee Clinical Campus, Wisconsi.
Long procedure time and somewhat suboptimal results hinder the widespread use of catheter ablation of complex arrhythmias such as atrial fibrillation (AF). Due to lack of contrast differentiation between the area of interest and surrounding structures in a moving organ like heart, there is a lack of proper intraprocedural guidance using current imaging techniques for ablation. Cardiac image registration is currently under investigation and is in clinical use for AF ablation. Cardiac image registration, which involves integration of two images in the context of the left atrium (LA), is intermodal, with the acquired image and the real-time reference image residing in different image spaces, and involves optimization, where one image space is transformed into the other. Unlike rigid body registration, cardiac image registration is unique and challenging due to cardiac motion during the cardiac cycle and due to respiration. This review addresses the basic principles of the emerging technique of registration and the inherent limitations as they relate to cardiac imaging and registration.
Key Words : Atrial Fibrillation, Imaging, Registration.
Correspondence to: Dr. Jasbir Sra, M.D., F.A.C.C., 2801 W. Kinnickinnic River Pkwy #777, Milwaukee, WI 53215
414.649-3390
Fluoroscopy does not provide adequate anatomic visualization, due to poor soft tissue contrast, and fluoroscopically-guided intracardiac catheter-based procedures, such as AF ablation, are becoming increasingly complex. Improved intra procedural imaging of relevant anatomy would, therefore, aid in guiding the accurate and efficient placement of ablation lesions in complex anatomy such as the LA during AF ablation.
Three-dimensional (3D) imaging modalities such as computed tomography (CT) and magnetic resonance imaging (MRI) offer high quality anatomic visualization, given their excellent tissue contrast characteristics and high spatial and temporal resolution. Registration involves an optimal strategy of aligning different images.1,2 It provides a means for physicians to incorporate within a single view the varied information captured by different imaging modalities. Intrasubject intermodal registration using 3D imaging modalities is an area of current focus with regards to radiofrequency catheter ablation procedures.3-11 Cardiac motion of the heart and interpatient variability of anatomical features make the registration of cardiac images a unique challenge. Objective validation of the registration technique is, therefore, necessary before its utility as a tool for image-guided therapy can be properly assessed.
This review addresses the basic principles of registration and their impact on registration results and examines the validity of applying the registration process within the particular constraints of cardiac imaging. Although any modality can be used for registration, given the current interest in image-guided therapy for AF, LA-CT imaging and registration is the main focus of this review.
CT-MR Imaging and Segmentation Patients
Most medical images are in a digital format and are made up of a rectangular array of small square or rectangular elements called pixels. Each pixel provides the coordinate system of the image and an element in the image can be assessed by its two-dimensional (2D) position within this array. A typical CT slice, for example, is formed of 512x512 pixels and each corresponds to a portion of the cut through the patient measuring about 0.5x0.5 mm2.12 The matrix and the pixel size are related to the display field of view (FOV). If for example, the FOV is 25 cm, each pixel will be FOV/matrix size, 25/512 = 0.48 mm2. This dimension determines the limiting in-plane spatial resolution of the image. The 2D axial slices are then stacked together to form a 3D volume. Each pixel corresponds to a small volume element, called the voxel. The height of the voxel is determined by the slice thickness. If the axial slice thickness, in the above example, was 1.5 mm, the voxel size would be is 0.48 x 0.48 x 1.5 mm3. For MR brain imaging, typical voxels may be 0.9 x 0.9 x 3 to 5 mm3 with 256 x 256 pixels in a slice. It is also possible to acquire MR images with cubic voxels which can be used for registration.
In a CT imaging system configuration, an x-ray projects a fan-shaped beam that is collimated to lie within an X-Y plane of a Cartesian coordinate system, generally referred to as the imaging plane. The x-ray beam, after being attenuated by the organ through which it passes, impinges upon an array of radiation detectors. The intensity of the x-ray beam received at the detector array is dependent upon the attenuation of the x-ray beam by the object.
Pixel values are a measure of the x-ray attenuation in Hounsfield units (HU) where the HU= 1000- (4µ/µw-1), and µ is the average linear attenuation coefficient of the volume element represented by the pixel and µw is a linear coefficient of water for the effective energy at the beam exiting the patient. Thus water has an HU number of 0 and a region with a CT number of 100 HU has a linear attenuation coefficient that is 1% greater than the linear attenuation coefficient of water.
In the ECG-gated helical CT acquisition, the x-ray tube (and detector array) rotates continuously about the patient collecting the data, while the patient table (and the patient) is moved at a constant speed through the imaging plane, thus collecting a continuous sequence of consecutive axial images from a volume of patient’s anatomy. Faster scanning, as currently done with 64 slice CT scanners, allows a large volume of data in short periods of time. This allows 3D reconstruction of the image set resulting in true depiction and a more highly diagnostic anatomic image across the patient population.
The factors selected in scanning a patient include slice width, FOV, gantry rotation speed and volume of coverage as well as basic contrast injection protocols. Scan time is calculated from the scan volume (total distance traveled by the table) divided by the table speed. Also important is the pitch which is defined as ratio of distance the table moves per 360º rotation. A pitch of 1.0 thus means the patient table moves a distance of one slice. Similarly, a pitch of 0.2 means the gantry rotates 5 times as the table moves a distance equal to the collimator width.
To avoid respiration artifacts, scanning is performed during the breath held in inspiration or expiration. The acquired data is synchronized with the collection of the ECG (QRS) signal. The data acquired during consecutive cardiac time intervals can then be combined to produce an image of the heart at the same phase of the cardiac cycle throughout the volume. Retrospective gating allows alignment of images during any phase of the cardiac cycle due to the continuous helical acquisition.
A medical image standard known as DICOM (Digital Information and Communications in Medicine) is widely used. This allows data to be exchanged between scanners and viewing consoles. The American Radiological convention is to display axial images with the right side of the patient at the left and posterior side at the bottom when looking at the computer workstation screen.
MRI is based upon the principles of nuclear magnetic resonance, a spectroscopic technique to obtain microscopic chemical and physical information about molecules.12 MRI started out as a tomographic imaging technique, producing an image of the nuclear magnetic resonance signal in a thin slice through the human body. MRI has now advanced beyond a tomographic imaging technique to a volume imaging technique. Microscopic imaging is based on the absorption and emission of energy in the RF range of the electromagnetic spectrum. The human body is primarily fat and water. Fat and water possess hydrogen atoms. Hydrogen nuclei have an NMR signal which is imaged by MRI. Cardiac motion compensation is performed by synchronization of the image acquisition to the ECG signal. Both prospective and retrospective ECG gating can be performed. In addition to cardiac cycle motion, another source of image distortion is respiratory motion. Due to the development of faster MR imaging techniques, such as echo-planar imaging and turbo field echo imaging, it is possible to acquire images during short breath-hold of around 15 seconds. Recent technical advances have make it possible to acquire a stack of 12 slices, with around 25 cardiac phases each, in a single breath-hold of 20 seconds. As with the use of CT, PV anatomy can be clearly delineated using MRI.
LA Imaging and Segmentation
The process of dividing images into different regions to visualize areas of interest is called segmentatio.13-14 Because of issues such as spatial resolution, poor contrast, ill-defined boundaries, noise, or acquisition artifacts, segmentation is a difficult task. Image segmentation methods can be grouped into thresholding, boundary detection, and region identification.
Thresholding is the simplest but most effective segmentation method. During thresholding, pixels with intensities below a threshold value are assigned one class and the remaining pixels a different class. Regions are then formed by connecting adjacent pixels of the same class. Boundary edges are needed in various image analysis applications. Thus boundary extraction methods use information about intensity differences between adjacent regions to separate the regions from each other. If the intensities within a region vary gradually but differences of intensities between adjacent regions remain large, boundary detection methods will delineate the region. Region identification techniques then form regions by combining pixels of similar properties.
Detailed CT and MR studies15-17 have shown that anywhere from 65% to 80% of patients have 4 PVs, with left common and right middle seen in others. Part of the main trunk of the right superior PV passes immediately behind the right superior-SVC junction. It has also been shown that the right superior PV trunk branches out significantly sooner than do the left PVs. In 10% to 20% of cases, a middle PV arises independently on the right side.
The right inferior PV arises inferiorly and laterally to the right superior PV. It divides almost immediately, within 5 to 10 mm, into superior and inferior branches. The distance between the right superior and right inferior PVs across the canal ridge varies from 2 to 8 mm.
The left superior PV lies superior and posterior to the LA appendage. It enters the LA in a more vertical direction. It usually has multiple branches which ordinarily arise 10 to 20 mm from its base. The left inferior PV enters the LA more horizontally from a posterior and lateral position. It branches almost immediately. A common antrum of the left superior and inferior PVs is seen in 3% to 30% of patients. In a series of over 500 CT scans, in addition to the left common and right middle PVs
(Fig 1),18 we have seen other unusual anatomies including a common right PV, 3 PVs on the right and left, and a common ostium of the left inferior PVs
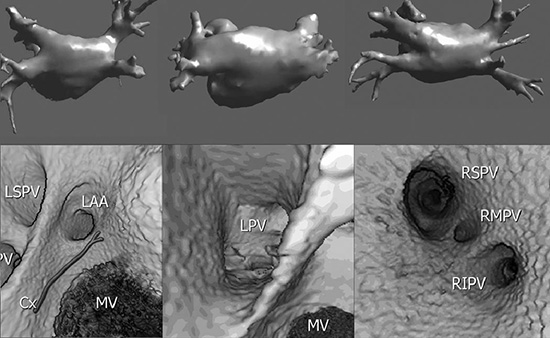
Figure 1. Three-dimensional (3D) and endocardial left atrial image reconstruction. Representative examples of 3D models of three different pulmonary vein morphologies, along with endocardial views, are depicted. As opposed to the standard four pulmonary veins in the left panel, the middle panel shows a common pulmonary vein and the right panel shows an additional right middle vein. The respective endocardial views, along with the mitral valve (MV), are shown in the bottom panel. The location of the circumflex artery (Cx) in the left panel is also shown. LAA = left atrial appendage; LIPV = left inferior pulmonary vein; LSPV = left superior pulmonary vein; RIPV = right inferior pulmonary vein; RMPV = right middle pulmonary vein; RSPV = right superior pulmonary vein. Reproduced with permission from Sra et al. J Cardiovasc Electrophysiol Images 2004; 15:247
Fundamentals of Image Registration
The image registration process entails combining images from two different image coordinate spaces, such that multiple anatomic features or points of interest are aligned.1,19,20 These points, called the fiducial points, are marked on the two image spaces. During the actual registration process, the image to be registered undergoes a mathematical transformation in all of its degrees of freedom, such that its image coordinate space is mapped onto that of the other image. For 3D structures, there are 6 degrees of positional freedom (translation along three orthogonal directions, and rotation along three axes). In addition, for the purpose of calibration, scaling factors between two image spaces must be accounted for in three dimensions. This registration process can be accomplished with a minimum of 3 non-collinear fiducial points matched in each of the image spaces. More points would increase the accuracy of the registration process. A translation factor can be achieved by using techniques and processes to align the center of the anatomical structures. Proper rotation can be calculated in terms of three angles (θx, θy, and θz). Assuming that the systems used for registration are calibrated such that the patient centered referential is the same in terms of orientation, there will be minimal rotation between the two systems. However, some rotational error may still occur due to movement of the heart and during respiration.
Several assumptions are generally made during this process. First, the structure undergoing registration is assumed to be a rigid body, which has undergone no changes or deformations from the time of imaging to the time of registration. This assumption may introduce limitations into the registration process when applied to a moving structure such as the heart. Furthermore, the final registration error is limited by how accurately fiducial points were identified in the respective image spaces.
Cardiac registration, in the context of the LA during AF ablation, is intermodal, with the acquired images and real-time reference image residing in different imaged spaces. Registration algorithms involve the optimization of a cost function by the choice of a transformation (T) which transforms one image space into the other. T can either be linear or nonlinear.19-21 Linear transformations are shape-preserving and are composed solely of rotations, translations, and isotropic scaling. Nonlinear transformations may deform both shape and size between images. A linear or nonlinear transformation decides the number of parameters involved in its definition. A linear transformation between 3D spaces is defined by 6 parameters (or degrees of freedom), where two positions of a rigid body can always be related to one another in terms of three translations and three rotations. As the voxel sizes in each image may not be similar for calibration purposes, three extra degrees of freedom, equating to scaling in each direction, are needed
(Fig. 2). A nonlinear transformation, as it may deform both shape and size, will require more degrees of freedom.
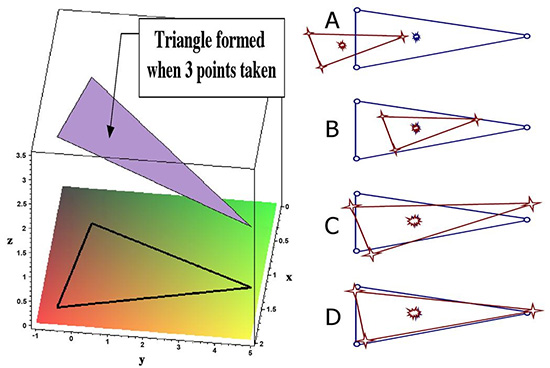
Figure 2. Simplified rigid body registration process. Left panel shows that once 3 fiducial points are taken a triangle is formed. In the right panel, A represents projection of CT (red) and reference image (blue). Panel B depicts initial step which involves translation of centroids. Panel C involves scaling to calibrate both images. Panel D shows rotation of the image to minimize the sum of the squared distances between fiducial points
The cost function is a measure of similarity (or dissimilarity) between the reference image and the transformed image and optimization is usually required to make the registration process accurate. Gradient descent method19 is used to calculate the cost function for the set of registration parameters described above. The direction of the greatest decrease of cost function can be mathematically computed at that set of parameters. A search is done in the direction of the greatest decrease until a new minimum is found. The parameter set is then updated to be the point at which the new minimum occurs. This process continues until minimal change occurs in the cost function with successful steps.
Registration methods fall into two broad categories, based on either geometric similarities between images or voxel intensity similarities. Voxel similarity-based methods have traditionally been used to assess serial changes in the brain using MR images and are not relevant to this review and will not be discussed further.
Geometry-based methods can be point-based or surface methods applied to both 3D-3D and 3D-2D registration models. Point-based registration methods depend upon the appropriate definition of two sets of corresponding fiducial points in each set of images to be registered. The cost function, the residual sum of squares (RSS) between the transformed points of the acquired image and their corresponding points in the real-time image, is determined and then applied to the entire acquired image data set. Thus, if 1N denotes the set of fiducial points in the acquired image, {y}1N denotes the corresponding points on the real-time image, i.e., the choice of T minimizes the error iN=1 IIt(x)-y1 II2. Fiducial point sets are either automatically determined based on external skin markers or manually determined based on anatomical landmarks. The former method is usually used in the registration of neuro images, as the static nature of the brain allows its internal organization to be characterized by its external features. The latter method has been used in cardiac image registration.20,21 For rigid structures such as the brain, three non collinear fiducial points may be sufficient to establish the transformation between the 3D image volumes. However, the larger the number of points used, the more any error occurring during collection of the points is averaged out. The algorithm for calculating the transformation between the 3D images involves first computing the average of the “centroids” of each set of points. The difference between the centroids in the 3D images tells us the translation that must be applied to one set of points. These points are then rotated about the new centroids until the sum of the squared distances between each corresponding point pair is minimized. The square root of the mean of this squared distance is referred to as fiducial registration error (FRE). The mathematical solution is known as the solution to the orthogonal Procrustes problem,22 named after a robber in Greek mythology who fitted his guests to a bed of the wrong size. The manual aspect of the fiducial point localization, however, has an inherent error, the fiducial localization error (FLE), associated with each point.
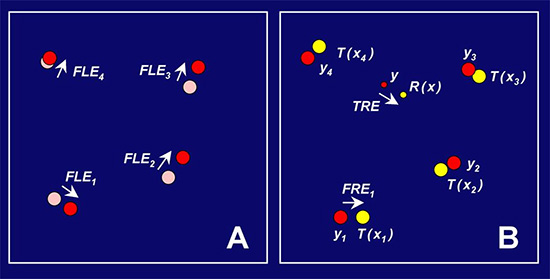
Figure 3. Schematic of point-based registration depicting errors. Panel A depicts the fiducial localization (FLE) error. Red circles represent positions at which the points are determined by the localization process in one of the two spaces involved in the registration process. The light circles represent the actual position. Panel B depicts point-based registration illustrating two measures of registration error. The red circles represent position y in one space. The lighter circles represent position x in the other space after they have been mapped by the registering transformation T. The larger, numbered circles are the points used to effect the registration. Fiducial registration error (FRE) is the alignment error between these. Target registration error (TRE) is the relevant registration error at a point (smaller circles) not used in the registration. Reproduced with permission from Sonka M, Fitzpatrick JM (eds). Handbook of Medical Imaging. V.II. Bellingham, WA:SPIE Press; 2000
Surface-based methods are used in registering a 3D image. Surface structure information is extracted from the CT or MR. Hence surface-based methods are mostly used in conjunction with these modalities. The process involves characterizing the surface represented in each of the images to be registered then deriving the best transformation T between them. The cost function here is a display function which provides a measure of distance between the surfaces. One of the surfaces is represented by a set of discrete points while the other has a surface representation. Surface can be represented as faceted surfaces, implicit surfaces or parameterized surfaces. Initially, a correspondence between the point set and the surface must be chosen. T is then optimized to minimize registration errors. One problem with the method is that, although T is not rigid in the context of cardiac registration, to achieve computational efficiency with this method, it is often assumed to be rigid. Another potential problem with this technique is that it is highly dependent upon identification of corresponding surfaces, yet different imaging modalities can provide very different image contrast between corresponding structures.
Registering 3D and 2D Images
FIf the geometry of the image acquisition is tightly controlled, 2D images may be registered purely via a rotation and two orthogonal translations. Clinically relevant examples of this are rare. 3D to 3D image registering using CT or MR is becoming more widespread. The assumption is usually made that the internal anatomy of the patient has not distorted or changed in spatial relationship between organs so that the imaged part of the body behaves as a rigid body. In this case, three translations and three rotations will bring the image into registration as described before. The rigid transformations, a subset of linear transformations, are composed solely of rotations and translations. If T is rigid between images of the same dimensions, the use of the singular value decompositions (SVD) method has been determined to provide T with a good degree of accuracy and stability, and is favorable for its noniterative time efficiency.23 SVD is a technique for expressing a given matrix as a product of three component matrices which reveal structural information about the original matrix. Orthogonal matrices are square matrices with row and column vectors directionally perpendicular to each other, having measurable magnitudes. Thus, if T is rigid, it can be expressed as follows: T (x) = (x) + t where is an orthogonal rotation matrix and t is a translational vector.
The process of registration becomes more complicated if one of the images represents a projection of physical space, as is the case with most optical images and conventional x-ray radiographs. These images are called “projection” images. One point in an x-ray image will correspond to some combination of the x-ray attenuation value along the line in the patient leading from the x-ray source to the imaging plane. This means that one point in the radiograph will correspond to a line of points through a CT or MR image. In optical images, only the visible surface will contribute to the image.
When T is constrained to be a perspective projection transformation from a 3D space to a 2D space, the problem is referred to as Projection Procrustes Problem.22 If, for example, x1 is a matrix of image points in a 3D image space, x2 a matrix of corresponding points in the 2D image space and a projection transformation, a solution of the Projection Procrustes Problem minimizes II x1 - x2 II. This problem is iteratively solvable when x1 and x2 are known; a non iterative solution based upon the SVD technique described before can also be found in a similar manner to that outlined before.
For computational efficiency, transformations are restricted to scaling, translations and rigid rotations in the x-ray space, and are based on the optimized alignment of corresponding features or other image properties. The details may vary, but they are described mathematically as  (af (x) +t) » y where f is a choice function that selects an appropriate 2D slide of the CT volume. Based on the orientation of the x-ray C-arm, a is a scaling parameter that standardizes the sizes of the images, t is a 2D translational parameter that aligns the centers of the images and  is a rotational matrix that completes the alignment process. In the CT x-ray registration, a, t , and  are the usual parameters involved; however, the choice function f can also be thought of as an additional parameter. Registrations that are non rigid and incorporate more parameters generally produce more accurate results. However, as results for interventional purposes need to be obtained in real time, a compromise has to be reached in registration design between accuracy and quick computational design. Surface-based methods are used in registering a 3D image. Surface structure information is extracted from the CT or MR. Hence surface-based methods are mostly used in conjunction with these modalities. The process involves characterizing the surfaces represented in each of the images to be registered then deriving the best transformation T between them. The cost function here is a display function which provides a measure of distance between the surfaces. One of the surfaces is represented by a set of discrete points while the other has a surface representation. Surfaces can be represented as faceted surfaces, implicit surfaces or parameterized surfaces. Initially, a correspondence between the point set and the surface must be chosen. T is then optimized by some form of the gradient descent technique described before. The optimization algorithm selects T to minimize the sum of weighted distances in the disparity function between the reference image surface and the set of transformed points and it is known as the iterative closest-point algorithm.
One problem with the method is that, although T is not rigid in the context of cardiac registration, to achieve computational efficiency with this method, it is often assumed to be rigid. Another potential problem with this technique is that it is highly dependent upon identification of corresponding surfaces, yet different imaging modalities can provide very different image contrast between corresponding structures. Currently, computer-assisted segmentation almost always requires some manual editing and adjustment.
Clinical Applications for (LA) Registration
Many studies have been performed recently to implement semi-automated 3D-3D and 3D-2D registration strategies. This section describes some recent advances made in the registration of acquired, structurally-revealing 3D images with real-time images. Table 1 depicts different registration techniques for the LA and various errors reported with some of these studies.
Table 1. Registration Techniques and Errors
| Author |
Ref |
Registration Technique |
Pts./other |
Error mm |
Comments |
| Sra |
HR, 2005 |
CT/ESI |
Animal Model |
2.0±3.6 |
target registration error |
| Tops |
HR, 2005 |
CT/CARTO |
16 |
1.8±1 |
feasability- mapping points and CT surface |
| Dong |
JCE, 2006 |
CT/CARTO |
16 (8 CT) |
3.0±0.4 |
technique as above |
| Malchano |
JCE, 2006 |
CT/CARTO |
13 |
12±11 (insp)
4.7±0.9 (exp)
|
technique as above |
| Kistler |
JCE, 2006 |
CT/CARTO |
47 |
2.4±0.4 |
technique as above-CartoMerge (83% vs. 60%) |
| Dong |
Circ, 2006 |
CT/CARTO |
Animal Model |
1.8±1.0 |
target registration error |
| Fahmy |
JCE, 2007 |
CT/CARTO |
124 |
5.6±3.2 (post)
9.1±2.5 (ant)
|
ICE-guided assessment |
| Zhong |
HR, 2007 |
CT/CARTO |
16 |
16±12 (right)
11±7 (left)
|
ICE-guided assessment |
| Sra |
Circ, 2006 |
CT/fluoro |
Phantom, 20 Pts |
1.4±0.5 |
feasability |
| Sra |
JCE, 2007 |
CT/fluoro |
50 |
--- |
outcome study (84% vs. 67%) |
3D Anatomical Mapping Systems
Anatomical mapping systems provide the 3D position of a navigational catheter within the cardiac chamber of interest and, in some instances, can also be used to construct 3D maps of the cardiac chamber. Magellan and CARTO (Biosense Webster, Inc., Diamond Bar, CA, USA).24,2 use the electromagnetic position of the catheter tip, based on an electromagnetic locator pad which is placed below the patient, and a reference catheter at a fixed external (usually posterior) location. LocaLisa (Medtronic, Inc., Minneapolis, MN, USA) and NavX (St. Jude’s Medical. Inc., Minneapolis, MN, USA) use voltage gradients generated by external electrical fields to spatially orient and localize the catheter tip.26,27 EnSite balloon (St. Jude’s Medical, Inc., Minneapolis, MN, USA)28 uses an electrically-coded catheter and a multielectrode mapping balloon to create maps and define the location of the navigational catheter.
Point and Surface-Based Registration
In one of the earlier studies, point-based registration was performed using CT images and the EnSite system.4 Fiducial points (≥ 3 non collinear points) were manually inserted in the form of pacing leads into the LA of a canine model and CT images were recorded. Following segmentation of the LA, a wire-frame model of the LA was imported to the noncontact, multielectrode array-based EnSite mapping system. Goodness-of-fit was evaluated qualitatively by observing a cloud of points post registration. When registration was tested by navigating the catheter to the buried electrodes and delivering radiofrequency pulses at these points, the mean fiducial registration error was 6.9±2.2 mm and the target registration error was 2.0±3.6 mm.
A study by Solomon et al25 attempted the registration of porcine cardiac CT images with the real-time catheter location provided by the Magellan mapping system. A point-based registration method was used with fiducial points determined by external markers placed on the swine’s chest prior to CT imaging. Accuracy of the registration was determined by position error, distance between the actual location of the marker and the location where the navigational system registered the point (fiducial registration error). The accuracy was determined to be within 4.69±1.70 mm. A reference catheter taped to the swine’s chest indicated that lateral motion due to respiration accounted for increased inaccuracy.
Reddy et al3 performed a similar study on the registration of MR images using the CARTO system. The main difference was that a surface-based method was chosen for registration in which the MR images were created and the points were marked out on the left ventricle and aorta via the CARTO. An iterative closest-point algorithm was used to register the image. The results indicated that registration of the left ventricle alone resulted in inaccurate alignment. Inclusion of the aorta in the registration process rectified this error.
A clinical application of this technique is now available which uses the CARTO system and either CT or MR (CartoMerge™).5,8-11 A combination of landmark and surface registration is used to register CT or MR with the CARTO system. Initially, several landmarks, usually 3, are manually chosen and annotated. Following this, the 3D LA reconstructed image from the CT or MR is superimposed upon the electroanatomic map created by the CARTO mapping system
(Fig. 4).
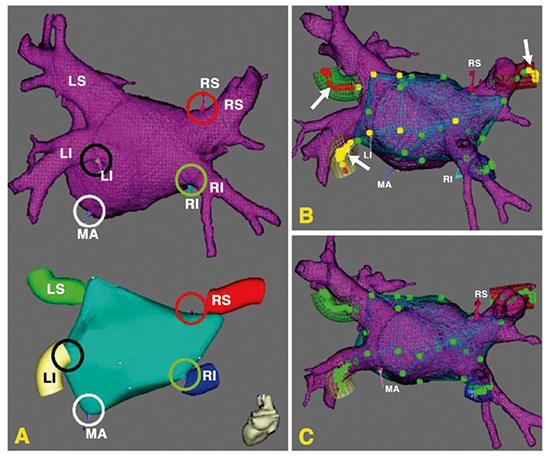
Figure 4. Left atrium (LA) registration. A: Landmark pairs (highlighted with colored circles) at the 6 o’clock mitral annulus (MA) position and the junctions of the LA and right superior pulmonary vein (RS), right inferior pulmonary vein (RI), left inferior pulmonary vein (LI) were annotated on the 3D CT LA surface reconstruction (upper image, shown as wire-frame) and the LA electroanatomic map (lower image, shown as solid shell) with colored tubes representing pulmonary veins (PVs). B: After landmark registration, the 3D LA surface reconstruction was superimposed on the electroanatomic map (shown as mesh). Note, the misalignment of the LS, LI, and RS between the two image datasets, indicated by the yellow or red color of the PV points (arrows) sampled in those PVs. C: After surface registration was executed, the PV alignment between the two image datasets was significantly improved, indicated by the green color of the PV points. The green, yellow, and red colors of the electroanatomic map points indicate their distance of <5 mm, 5–10 mm, and >10 mm, respectively, from the registered CT reconstruction surface. Reproduced with permission from Dong et al. J Cardiovasc Electrophysiol 2006;17:459-466
In a feasibility study,5,16 patients undergoing AF ablation underwent fusion of LA images obtained from a multislice CT scan with an electroanatomic mapping system using the CartoMerge™. A detailed electroanatomic map of the LA was created first. The 3D CT image and the electroanatomic map were displayed simultaneously on the CARTO screen. Using fluoroscopy and intracardiac echocardiography guidance, a landmark was then placed on the electroanatomic map and on the CT image on the same spot. Next, the surface registration algorithm was implemented. This algorithm composed the best fit of the two structures based on minimizing the distance between the two landmarks and the distance between all mapping points and the CT image. Following this, it was shown that it was feasible to perform registration guided therapy.
A main concern in this and prior studies was that they assumed the transformation between the CT or MR and the interventional system was rigid in nature and thus did not take into account cardiac motion errors.
Several recent studies have tested the accuracy of CT-MR registration. Dong et al,8 attached 2.3 mm CT fiducial markers to the epicardial surface of each cardiac chamber in 9 mongrel dogs. CT images were registered to the electroanatomic maps of each cardiac chamber. To assess accuracy, targeted ablations were performed at each of the fiducial markers guided only by the reconstructed images. At autopsy, the position error was 1.92±0.9 mm for the right atrium, and 1.8±1.0 mm for the LA. Retrospective analysis revealed that a combination of landmark registration and the target chamber surface registration resulted in < 3mm accuracy in all chambers.
However, other clinical studies have shown significant errors during registration using these techniques. One study showed poor registration during inspiration but good alignment during expiration.11 In a study by Fahmy et al29,124 patients underwent AF ablation guided by registration of images of the left atria created using CartoMerge™. Different fiducial landmarks were selected and spatial accuracy was further validated using intracardiac echocardiography. Surface registration was then performed and its impact on integration using the fiducial markers was assessed. The best landmark registration was achieved when the posterior points on the PVs were selected (5.6±3.2). Landmarks taken on the anterior wall, the LAl appendage or the coronary sinus resulted in greater registration error (9.1±2.5). As opposed to the previous study, in this study, surface registration resulted in shifting of the initially registered landmarks leading to a more significant level of error (from 5.6±3.2 to 9.2±2.1 mm; 95% CI 4.2-3.05).
Another recent study measured the accuracy of CartoMerge™ in guiding posterior LA ablation.30 In this study in 16 patients undergoing AF ablation, CT images of the LA were merged with CARTO. Encircling of the left and right PV vestibules was then performed, guided solely by intracardiac echocardiography, with point locations saved on a CartoMerge™ image to which the operator was blinded. The accuracy of the CartoMerge™ image was then assessed by measuring the distance from the location of each ablation point on the image to its actual anatomic location. A significant error of 1.6±1.2 cm in the right vestibule and 1.1±0.7 cm from the planned path of ablation was observed
(Fig. 5). Accuracy was improved when an end atrial contraction CT image was used.
Figure 5. CartoMerge image demonstrating a cut-away view of the right (right) and left (left panel) pulmonary vein region. Endocardial aspects are gray, and epicardial aspects are yellow. The actual lesion path is shown by the dotted line. Each red sphere demarcates an electroanatomic point taken along this path, which was acquired using ICE guidance while blinded to the CartoMerge image. Reprinted from Heart Rhythm, Vol. 17, Zhong et al. On the accuracy of CartoMerge for guiding posterior left atrial ablation in man, pages 595-60, 2007 with permission from Elsevier
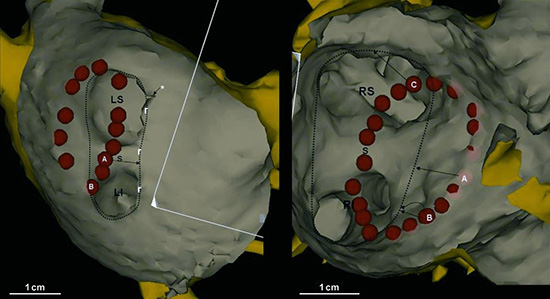
These reports suggest that these registration techniques have inherent limitations. Use of appropriate locations for fiducial points, cardiac cycle location and respiration could potentially help these approaches to registration.
Fluoroscopy-Based Registration
For low infrastructure and procedural costs, x-ray fluoroscopy remains the standard catheter navigation tool used in interventional laboratories. 3D-2D registration is much more complicated than the 3D-3D registration described before.
Registration of MR using biplane fluoroscopy and CT with single-plane fluoroscopy has been successfully performed.7,31,32 However, as only CT-fluoro integration has been tested in the context of AF ablation, this section discusses recent advances in fluoroscopy-based registration but focuses mainly on studies involving registration of 3D-LA models with 2D x-ray fluoroscopy.
In the study by Meyer et al,31 a registration algorithm was developed to register ultrasound-based 3D catheter coordinates, x1 as the fluoro view of the catheter with the corresponding coordinates given by the matrix x2. The study approximated the projection P, as described before, and then iteratively solved for the rigid transformation T such that II T ( x1)- x2 II was minimized. The study developed an initialization routine for T which resulted in a convergence of iterates in all the test cases. Four methods of error were defined and the method was tested using mathematical simulations and a multi-catheter phantom. The study concluded that use of a larger set of fiducial points along the catheter, with a wide spatial distribution, yielded registration results that were less sensitive to noise in the entries of x1 and x2.
The study by Rhode et al32 utilized a combined x-ray and MR interventional suite which produces both x-ray and MR images as separate outputs. The transformation that registered MR to x-ray images was presented as a product of point-based rigid transformations between each distinct space in the suite followed by a perspective projection transformation. The transformations were determined sequentially using a calibration object with ≥ 6 markers forming the set of fiducial points.
The registration was tested on a phantom and on post procedure patient images. A 2D target registration error was computed from the displacement of the MR transformed from the actual marker position on the x-ray fluoro view, representing the accuracy of registration. 3D target registration error was computed from 3D displacements between MR image markers and the reconstructed 3D positions of the markers on the x-ray image. The 3D reconstruction was done from the biplane x-ray images using the epipolar constraint. The 2D error for in vitro tests was 4.2 mm and the 3D error was 4.6 mm.
In the study by Ector et al,7 cardiac MR images were acquired in patients undergoing catheter ablation. Detailed 3D models of the LA were obtained. After contouring of endocardial cavities with cross-checking of different imaging planes, 3D models of the right atrium were merged with the contrast-filled right atrium seen on biplane fluoroscopy using custom-built software. The feasibility and accuracy of this merging process was determined in heart cast experiments and using electroanatomic mapping in patients. The reported error on heart casts was 0.2 ± 0.3 mm, and 1.9 to 2.5 mm in patients.
Fluoroscopic images are conic projections, with greater magnification of peripheral regions as compared to central ones. In contrast, CT images represent the synthesis of many x-ray projections obtained circumferentially around the object, thus essentially canceling the effect of this distortion. Assuming that the patient is positioned in the same position as when the CT scan was obtained, the height of various cardiac structures with respect to the table can be ascertained from the CT scan. Fortunately, the theoretical scaling of structures at each height above the table position is also well described with respect to the geometry of the fluoroscopy system (i.e. source and image-intensifier position). This information can then be used to scale and transform the CT image to match the fluoroscopic image space.
As fiducial selection (described in 3D to 3D image registration) in each of the image spaces introduces several challenges, we have instead used a coronary sinus catheter positioned through the superior vena cava as a fiducial marker and have used a transformation to align the coronary sinus catheter seen on fluoroscopy with the superior vena cava and the coronary sinus in the exported 3D model. In addition, for the purpose of scaling, the coronary sinus catheter is placed at a location identified by the intersection of a horizontal plane and the line joining the focal point of the fluoroscopic image. The horizontal plane is defined by the elevation above the anatomical reference system, the patient table. Once the location of the catheter and the actual and apparent dimensions of the elevation in the 3D model are known, the actual size of the same anatomical structure in the 3D and the fluoroscopic image can be determined. We recently described an implementation of a semi automated 3D-2D CT-fluoroscopy registration strategy.6 The accuracy of this system was found to be within 1.4 mm in phantom studies. This strategy was also assessed in patients undergoing AF ablation. Accurate registration was confirmed by: 1) PV angiography and 2) position and recordings on a 64-pole basket catheter located in the PVs. In a recent study33 patients with AF undergoing ablation were randomized to standard ablation techniques or ablation guided by the CT-fluoro registered images. Fluoroscopy time and total procedure time was significantly reduced in the CT-fluoro group (p<0.01) and there was a trend toward better outcome. It is also possible to mark ablation points on these images and synchronize the images to the cardiac cycle. This technique has been used in 154 patients and should be available for general use in the next 18 months.
As in the 3D-3D registration described before, the largest source of error with the 3D-2D registration strategy may be due to cardiac and respiratory motion. The complex translational, rotational, and conformational changes that occur with cardiac and respiratory motion will introduce error into the registration process when, as in 3D-3D registration, a static image is “aligned” with the real-time fluoroscopic image. Attempts at gating the registration process so that image integration occurred during the same phase of the cardiac cycle were successful in experiments conducted in our laboratory, thus fluoro images can be taken at the same phase as the segmented CT images, i.e., during diastole . Movement of the LA during respiration could potentially be eliminated by synchronizing registration to the respiratory cycle. In addition to cardiac gating, we have been able to take into account respiration gating and deposit ablation points on the surface of registered images as depicted in
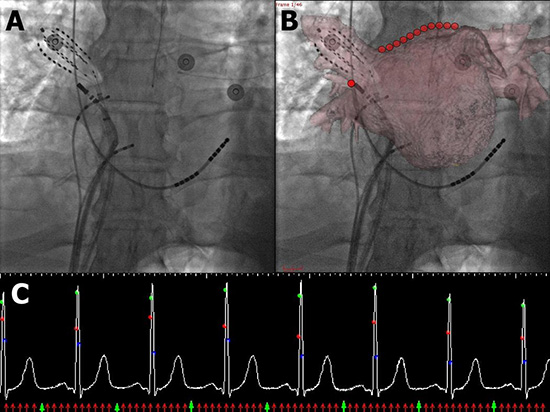
Figure 6.
Figure 6. ECG and respiration-gated registration. Fluoroscopic image is taken during the same cardiac cycle phase (75% as shown by green arrows), and respiration-gated (during inspiration) as CT imaging. Accurate registration is seen. System also allows marking of the ablation site (seen as red circles)
Ultrasound-Based Registration
Ultrasound-based registration has not been studied in detail, possibly due to difficulty in achieving automatic processing of these images in conjunction with images from other modalities.
Registration of 3D ultrasound has been attempted in the field of stress echocardiography34 where pre and post stress 3D ultrasound images were temporarily aligned and then spatially registered by optimization of mutual information between image voxel intensity sets, thus providing voxel intensity-based registration.
In the study by Packer et al,35 data on 5 patients was integrated from electroanatomic or noncontact mapping, CT anatomy, and intracardiac ultrasound. CT images of the LA were created. Ultrasound images of the PVs were generated using intracardiac, phased-array ultrasound with imaging at 5.5-10.0 MHz. A fused, CT ultrasound activation sequence was then constructed from an image set. On this framework, electroanatomic or noncontact mapping was demonstrated.
MR-Based Registration and Navigation
In a recent study, an electromagnetic catheter positioning system was superimposed on 3D MR images using fiducial markers].36 This allowed display of the catheter position on the anatomy obtained using the MR images in real time. In phantom models, the accuracy was 1.11±0.06 mm. Subsequently, in the swine model, in vivo accuracy was 2.74±0.52mm. Radiofrequency lesions were repeated in the right atrium within an accuracy of 3.92 ±0.5 mm. This study suggests that ablation can be performed on the real-time images obtained using MR images. MR compatible catheters and the absence of 3D images may be some of the limitations which will need to be overcome.
Sources of Registration Error
The registration techniques described above afford a significant advantage over the less detailed geometry created by the currently available mapping systems, however, there are significant inherent limitations, as registration algorithms that assume rigid transformations between image spaces have been applied in many of these techniques. As described before, the initial fiducial points are picked up by the operator using fluoroscopy; the exact location of these points in the 3D space may be deceptive. As multiple points are needed for surface registration, the process takes more time, the manual component becomes more laborious. Given the dynamic motion of the heart and the large number of variables that influence its movement and shape, careful validation of any technique used for cardiac image registration is important.
This section explores the unique sources of error that make cardiac image registration challenging. An understanding of the potential errors inherent in the registration technique is essential to create error-free registration.
Errors common to intermodal registration
In the registration algorithms, the voxel and pixel dimensions have to be accurately reported by the imaging system. CT imaging systems, for example, often produce images where the thickness of reconstructed 2D slides are different from the voxel height and width, due to the somewhat limited resolution capabilities of the system along the z axis.
In 3D to 3D registration, this limited resolution capability has been shown to cause anisotropic error measurement in three dimensions, with the error echoing the reported voxel dimensions.37 A sub pixel/voxel measurement could eliminate these errors. Standardizing the size of images involves approximate measurements which can potentially introduce errors into the registration process. Since the actual position and size of the heart may not be known, approximate measurements of the expected distances and heights is sometimes used.38 The ratios of the actual versus the apparent diameters of ablation catheters in the registered image can be considered to help determine scale.
As described before, the x-ray image is a conic projection with greater magnification present in the peripheral regions of the image than in the central region. The 3D CT image, on the other hand, does not have this nonisotropic magnification. Thus, if error is present in the computation of scale, its effect would be amplified throughout the registration process.
Fiducial Point/Feature Selection
Location of fiducial points needs to be accurate. However, as these points are usually taken using 2D fluoroscopy, there are inherent limitations in using fiducial points selected based on fluoroscopy alone. As demonstrated in the prior study,29 fiducial points taken in posterior PV locations seem to give the best registration results. It is also important to take at least 3 non collinear points to get the best results. As discussed before, resolution differences also exist between CT and other 3D or 2D images.
Manual interaction is needed for feature segmentation or point localization, which can be associated with the well-documented fiducial localization error.19 Some choices of cost and optimization functions are more sensitive to accuracy of fiducial points. Registration based on alignment of features that may not represent identical regions is thus a potential source of error.
Studies have shown that accuracy is sensitive to the potential accuracy of spatial distribution of fiducial points. Wide asymmetric distribution of fiducial points produces more accurate registration. Similarly, regarding the selection of fiducial features, registration based solely on alignment of features that may not represent identical regions is also a potential source of error.
Errors Specific to Cardiac Image Registration
In addition to the errors described before, the possible sources of error specific to cardiac image registration are: 1) inconsistency in patient positioning, 2) cardiac gating during CT acquisition, 3) cardiac chamber movement in various stages of the cardiac cycle, and 4) cardiac chamber movement due to respiration.
Efforts should be made to align the patient identically on each imaging table to help ensure same orientation during both imaging modalities. Some variability in patient alignment and posture37 can result in a corresponding variability in the orientation of the imaged cardiac chamber. Factors that influence this variability include differences in imaging table surfaces, such as curvature and cushioning. Additionally, it is impossible to guarantee that the patient is in exactly the same position since there is usually a significant time-lapse between acquisitions.
The orientation of the cardiac chamber changes with the orientation of the body. For this reason, this type of interscan movement can be described as a rigid body rotation of the heart with the spine. A change in the interscan angle of 5 degrees, for example, may give a rotational error of 0.44 cm in a 5 cm organ with a field-of-view of about 20 cm. Thus it is important that any changes in interscan imaging are taken into account and minimized.
Cardiac and Respiration Gating
In addition to interscan movement resulting in different orientations of the imaged chamber, movement of the object between the same acquisitions can cause errors, as it is not possible to physically arrest the heart. In the cardiac chamber, this issue is amplified several-fold due to continuity and extent of movement of the heart. The cardiac deformities caused by non rigid cardiac motion have been broken down into as many as nine different modes.
Correction for some of these problems during the CT gating technique38 involves the patient holding breath for several heartbeats while 2D axial slices are sequentially obtained at fixed points of the cardiac cycle. These slices are stacked together and interpolated to create a smooth, static 3D CT image. Although the likelihood that the heart returns to a fixed shape at fixed points of the cardiac cycle is questionable, due to the variability in cardiac deformation,39 it is important that both the CT and the second image are ECG-gated to eliminate, as much as possible, errors due to cardiac motion. In the studies done in our lab, it was shown that the best images of the LA were obtained during segmentation of the LA at 75% of the cardiac cycle in sinus rhythm or at 45% during AF due to shorter R-R intervals.
Given the proximity of the heart to the diaphragm and given that the displacement of the diaphragm occurs in the direction of the heart, the effects of respiration are expected to be more pronounced. McLeish et al40 have determined that translations in the craniocaudal direction are the main types of movement seen in the cardiac chamber due to respiration. Less significant movements include translation and rotation in other directions. Noseworthy et al41 used MR angiography of the reconstructed LA and PVs at both end-expiration and end-inspiration. Their quantitative assessment of 3D images during respiratory phases revealed splaying of the PVs and reduction in the size of PV caliber in the right-sided PVs during held-inspiration. CT scanning can be done both during inspiration and now, due to rapid scanning techniques, during expiration. There is evidence based upon prior studies that respiration gating during expiration may be optimal.
The above findings do suggest that the effects of respiration need to be considered as their effect on target registration error may be amplified through the steps of the registration process.
Validation Strategies for Registration
Knowledge of the various sources of error is thus key to establishment of a gold standard since these errors will have to be accounted for in the simulation. An allowance for cardiac movement and respiration must be incorporated into the overall strategy, distinguishing cardiac image registration from other registration processes involving more static organs and structures. This could be accomplished either by distinguishing each component of a movement and introducing them into the simulation in combination or treating the movement as a single random process at a fixed point of the cardiac cycle. Animal studies can then be undertaken to complete the validation process.
Inadequate depiction of cardiac anatomy hinders the widespread application of fluoroscopy for AF ablation. Image registration involves correspondence between two images or an image and a physical space and provides a means for physicians to incorporate into a single view the varied information captured by different modalities which can then be used for the purpose of interventional planning and treatment. The deformations caused by cardiac chamber and respiration motion present unique problems for cardiac image registration. Although significant work has been done in the field of cardiac imaging, there is much room for the development of validation strategies for cardiac image registration. It is important that the issue of patient-specific movement of the cardiac chamber be quantified in evaluating registration algorithms. Proper application of the registration process, with careful adjustment for the constraints mentioned and use of the power of the computer to automate the process can greatly improve and simplify this technique.